Human Gene-Editing Trials: First U.S. Patients Treated With CRISPR
16 April 2019
https://www.npr.org/sections/health-shots/2019/04/16/712402435/first-u-s-patients-treated-with-crispr-as-gene-editing-human-trials-get-underway
Locus is one of several companies that are trying to use CRISPR to fight health problems by targeting only bad bacteria in the body and leaving the good ones alone. Locus scientists have created a cocktail of three phages that have been modified using CRISPR, which was discovered by studying the immune systems of bacteria. "What we've learned how to do is reprogram that immune system to attack itself," says Paul Garofolo, the company's CEO. "We load the viruses up with CRISPR constructs, which essentially work like little Pac-Men. They go into a target bacteria cell, and they chew up the DNA of that target. It makes them much more potent killers."
For an excellent History of Bacteriophages visit http://www.intralytix.com/index.php?page=hist
Directed Evolution of a Mycobacteriophage
25 April 2019. by María Cebriá-Mendoza,Rafael Sanjuán and Pilar Domingo-Calap Antibiotics 2019, 8(2), 46; https://doi.org/10.3390/antibiotics8020046
Abstract Bacteriophages represent an alternative strategy to combat pathogenic bacteria. Currently, Mycobacterium tuberculosis infections constitute a major public health problem due to extensive antibiotic resistance in some strains. Using a non-pathogenic species of the same genus as an experimental model, Mycobacterium smegmatis, here we have set up a basic methodology for mycobacteriophage growth and we have explored directed evolution as a tool for increasing phage infectivity and lytic activity. We demonstrate mycobacteriophage adaptation to its host under different conditions. Directed evolution could be used for the development of future phage therapy applications against mycobacteria. Full article
Efficacy & potential of phage therapy against multidrug resistant Shigella spp
Efficacy and potential of phage therapy against multidrug resistant Shigella spp.
Swee-Seong Tang, Sudhangshu Kumar Biswas, Wen Siang Tan, Ananda Kumar Saha, Bey-Fen Leo
Pub.5th April 2019. PeerJ Tang et al. (2019), PeerJ, DOI 10.7717/peerj.6225
This review surveys the extent of outbreaks of shigellosis and their effects, and then investigates the treatments of Shigella spp. using both antibiotic and phage therapy. It contains a chronological description of the emergence of Shigella spp. as a multidrug resistant pathogen, as well as outlining the limitations of antibiotics against multidrug resistant bacterial strains.
Conclusion: Phage therapies for Shigella spp. and other pathogenic bacteria have been studied and applied for about a century, but phage therapy as an antibacterial treatment in general has not received much attention due to lack of clinical knowledge and public awareness of phages. However, given that the development of novel antibiotics is laborious, time consuming and costly, it makes eminent sense to seek alternative antimicrobial approaches to combat drug resistant pathogens. While it inevitably has some drawbacks, phagebased biocontrol and bacteriophage therapy are very promising approaches to combat the challenge of pathogenic bacterial infections, particularly when the search for new antibiotics is stagnating. Moreover, phages have many unexploited potentials as an alternative to antibiotics, both due to the range of intrinsic variation in their mode of action, also due to almost unlimited variety of phages and their ability to evolve in situ to successfully deal with bacterial resistance.
The potential of phage therapy has been acknowledged and revisited by many scientists over the last few decades, and there has been a rejuvenation of research into phage therapy.
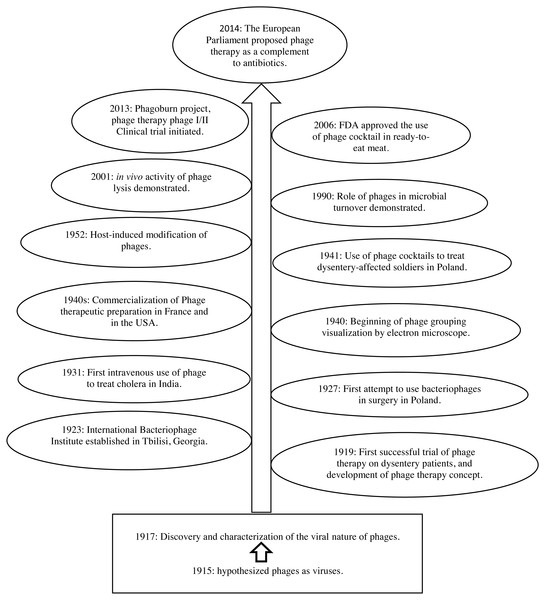
Figure 1: Milestones in phage therapy research, adapted from Elbreki et al. (2014) and Salmond & Fineran (2015). Download full-size image DOI: 10.7717/peerj.6225/fig-1
The FDA has approved bacteriophages as GRAS and allowed the application of phages as food additives in 2006, which is a significant boost to phage therapy research. Nevertheless, the therapeutic application of phages still requires extensive studies, judiciously performed clinical trials, and importantly well-defined regulatory guidelines.
Currently, phage therapy is encouraged in many parts of the world because policymakers consider growing MDR as a serious health problem. This awareness should further encourage researchers to study the biological properties of phages, which eventually increases their safety and efficacy. Furthermore, genetically modified phages could help to solve the issues of patent filing and as a result increase the interest of pharmaceutical and biotechnology companies to produce phage-products. Finally, cocktails of natural phages and genetically modified phages could open new perspectives for successful phage therapy in the future, particularly against the major challenge of Shigella and Shigella-like multidrug resistant bacteria.
Submitted 12 October 2017 Accepted 4 December 2018 Published 5 April 2019 Corresponding author Swee-Seong Tang, sstang [at] um.edu.my, sstang2107 [at] hotmail.com Academic editor Mark Enright Additional Information and Declarations can be found on page 18 DOI 10.7717/peerj.6225 Copyright 2019 Tang et al. Distributed under Creative Commons CC-BY 4.0 OPEN ACCESS
Environmental structure drives resistance & antibodies during phage therapy
Environmental structure drives resistance to phages and antibiotics during phage therapy and to invading lysogens during colonisation
Pub. online 28 February 2019. JorgeA. Moura de Sousa & Eduardo P. C. Rocha.
www.nature.com/scientificreports
Microbial communities are shaped by bacteriophages through predation and lysogeny. A better understanding of the interactions between these processes across different types of environments is key to elucidate how phages mediate microbial competition and to design efficient phage therapies. We introduce an individual-based model (eVIVALDI) to investigate the role of environmental structure in the elimination of a population with a combined treatment of antibiotics and virulent phages, and in the invasion of a population of phage-sensitive bacteria by lysogens. We show that structured environments facilitate the emergence of double resistance, to antibiotics and phages, due to limited diffusion of phage particles and increased nutrient availability from dead cells. They also hinder phage amplification, thus decreasing the generation of phage genetic diversity and increasing the unpredictability of phage-bacteria arms-races. We used a machine learning approach to determine the variables most important for the invasion of sensitive populations by lysogens. They revealed that phage-associated traits and environmental structure are the key drivers of the process. Structured environments hinder invasions, and accounting for their existence improves the ft of the model to published in vivo experimental data. Our results underline environmental structure as key to understand in vivo phage-bacteria interactions.
Microbial Evolutionary Genomics, Institut Pasteur, CNRS, UMR3525, Paris, 75015, France. Correspondence and requests for materials should be addressed to J.A.M.d.S. (email: jorge-andre.sousa [at] pasteur.fr)
Bacteriophages: Protagonists of a Post-Antibiotic Era
Bacteriophages: Protagonists of a Post-Antibiotic Era
2018 Antibiotics. Bacteriophages: Protagonists of a Post-Antibiotic Era
Pilar Domingo-Calap, Jennifer Delgado-Martínez
Despite their long success for more than half a century, antibiotics are currently under the spotlight due to the emergence of multidrug-resistant bacteria. The development of new alternative treatments is of particular interest in the fight against bacterial resistance. Bacteriophages (phages) are natural killers of bacteria and are an excellent tool due to their specificity and ecological safety. Here, we highlight some of their advantages and drawbacks as potential therapeutic agents. Interestingly, phages are not only attractive from a clinical point of view, but other areas, such as agriculture, food control, or industry, are also areas for their potential application. Therefore, we propose phages as a real alternative to current antibiotics.
Monash biologist awarded over $130,000 to investigate bacteriophage therapy
https://www.monash.edu/science/news/current/monash-biologist-awarded-over-$130,000-to-investigate-bacteriophage-therapy
02 November 2018. Research on bacteriophages has won Monash University biologist Dr Jeremy J. Barr, a Ramaciotti Award for Biomedical Research with a grant of $137,534. The Clive and Vera Ramaciotti Foundation, this week announced $784,731 in funding to seven biomedical researchers.
Dr Barr is the Head of the Bacteriophage Biology Research Group at the Monash University School of Biological Sciences. Dr Barr's lab studies bacteriophages -- specialist viruses that only infect and kill bacteria -- and investigates their role and function within the human body.
Bacteriophage Procurement for Therapeutic Purposes
Bacteriophage Procurement for Therapeutic Purposes
2016 Published in Front. Microbiol. DOI:10.3389/fmicb.2016.01177
Beata Weber-Dąbrowska, Ewa Jończyk-Matysiak, +3 authors Andrzej Górski
Bacteriophages (phages), discovered 100 years ago, are able to infect and destroy only bacterial cells. In the current crisis of antibiotic efficacy, phage therapy is considered as a supplementary or even alternative therapeutic approach. Evolution of multidrug-resistant and pandrug-resistant bacterial strains poses a real threat, so it is extremely important to have the possibility to isolate new phages for therapeutic purposes. Our phage laboratory and therapy center has extensive experience with phage isolation, characterization, and therapeutic application. In this article we present current progress in bacteriophages isolation and use for therapeutic purposes, our experience in this field and its practical implications for phage therapy. We attempt to summarize the state of the art: properties of phages, the methods for their isolation, criteria of phage selection for therapeutic purposes and limitations of their use. Perspectives for the use of genetically engineered phages to specifically target bacterial virulence-associated genes are also briefly presented.
HostPhinder: A Phage Host Prediction Tool
HostPhinder: A Phage Host Prediction Tool
2016 Published in Viruses HostPhinder: A Phage Host Prediction Tool
DOI:10.3390/v8050116 Julia Villarroel, Kortine Annina Kleinheinz, +4 authors Mette V. Larsen
The current dramatic increase of antibiotic resistant bacteria has revitalised the interest in bacteriophages as alternative antibacterial treatment. Meanwhile, the development of bioinformatics methods for analysing genomic data places high-throughput approaches for phage characterization within reach. Here, we present HostPhinder, a tool aimed at predicting the bacterial host of phages by examining the phage genome sequence. Using a reference database of 2196 phages with known hosts, HostPhinder predicts the host species of a query phage as the host of the most genomically similar reference phages. As a measure of genomic similarity the number of co-occurring k-mers (DNA sequences of length k) is used. Using an independent evaluation set, HostPhinder was able to correctly predict host genus and species for 81% and 74% of the phages respectively, giving predictions for more phages than BLAST and significantly outperforming BLAST on phages for which both had predictions. HostPhinder predictions on phage draft genomes from the INTESTI phage cocktail corresponded well with the advertised targets of the cocktail. Our study indicates that for most phages genomic similarity correlates well with related bacterial hosts. HostPhinder is available as an interactive web service [1] and as a stand alone download from the Docker registry [2]
